# point_labeler
**Repository Path**: zzj110/point_labeler
## Basic Information
- **Project Name**: point_labeler
- **Description**: No description available
- **Primary Language**: Unknown
- **License**: MIT
- **Default Branch**: master
- **Homepage**: None
- **GVP Project**: No
## Statistics
- **Stars**: 0
- **Forks**: 0
- **Created**: 2020-10-04
- **Last Updated**: 2020-12-19
## Categories & Tags
**Categories**: Uncategorized
**Tags**: None
## README
# Point Cloud Labeling Tool
Tool for labeling of a single point clouds or a stream of point clouds.
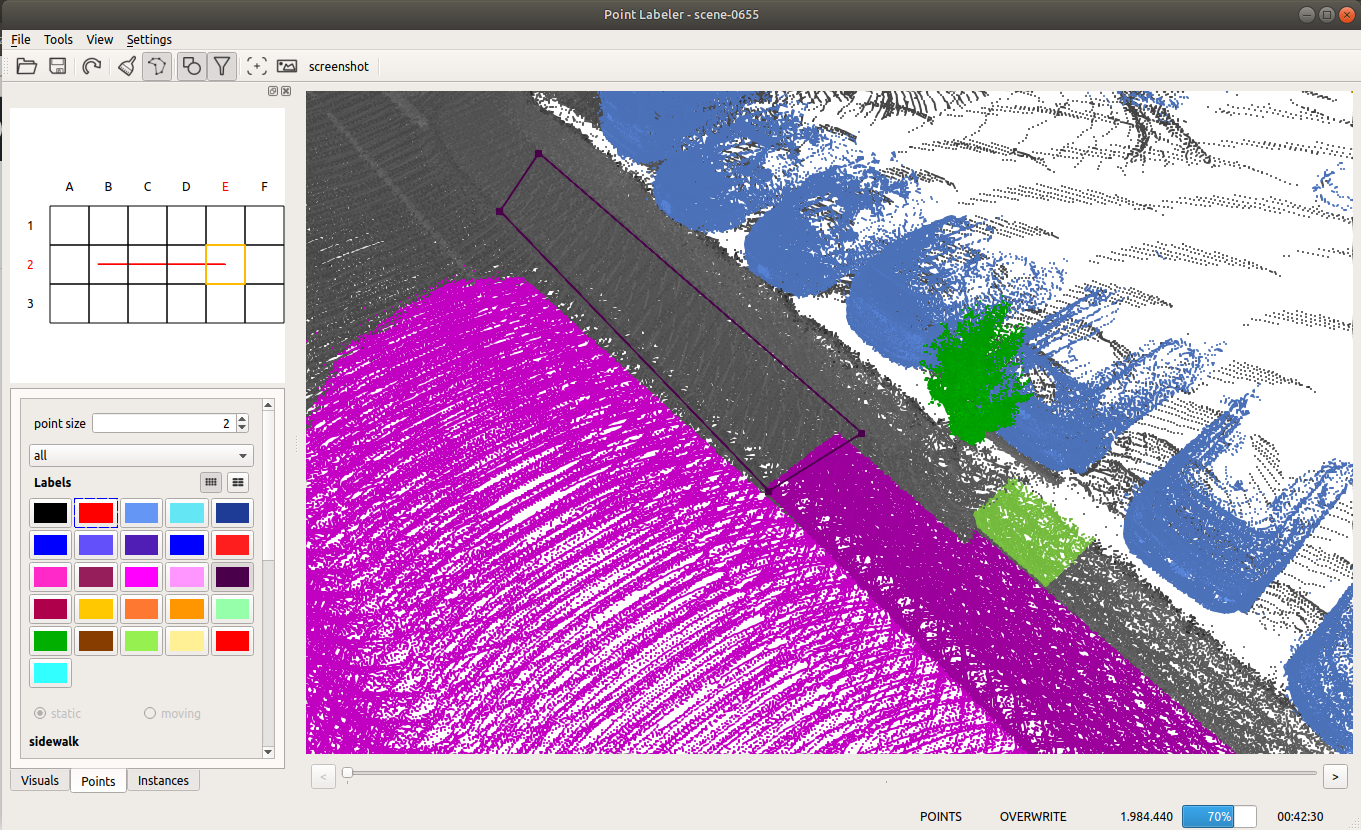 Given the poses of a KITTI point cloud dataset, we load tiles of overlapping point clouds. Thus, multiple point clouds are labeled at once in a certain area.
## Features
- Support for KITTI Vision Benchmark Point Clouds.
- Human-readable label description files in xml allow to define label names, ids, and colors.
- Modern OpenGL shaders for rendering of even millions of points.
- Tools for labeling of individual points and polygons.
- Filtering of labels makes it easy to label even complicated structures with ease.
## Dependencies
* catkin
* Eigen >= 3.2
* boost >= 1.54
* QT >= 5.2
* OpenGL Core Profile >= 4.0
* [glow](https://github.com/jbehley/glow) (catkin package)
## Build
On Ubuntu 16.04 and 18.04, most of the dependencies can be installed from the package manager:
```bash
sudo apt install git libeigen3-dev libboost-all-dev qtbase5-dev libglew-dev catkin
```
Additionally, make sure you have [catkin-tools](https://catkin-tools.readthedocs.io/en/latest/) and the [fetch](https://github.com/Photogrammetry-Robotics-Bonn/catkin_tools_fetch) verb installed:
```bash
sudo apt install python-pip
sudo pip install catkin_tools catkin_tools_fetch empy
```
If you do not have a catkin workspace already, create one:
```bash
cd
mkdir catkin_ws
cd catkin_ws
mkdir src
catkin init
cd src
git clone https://github.com/ros/catkin.git
```
Clone the repository in your catkin workspace:
```bash
cd ~/catkin_ws/src
git clone https://github.com/jbehley/point_labeler.git
```
Download the additional dependencies:
```bash
catkin deps fetch
```
Then, build the project:
```bash
catkin build point_labeler
```
Now the project root directory (e.g. `~/catkin_ws/src/point_labeler`) should contain a `bin` directory containing the labeler.
## Usage
In the `bin` directory, just run `./labeler` to start the labeling tool.
The labeling tool allows to label a sequence of point clouds in a tile-based fashion, i.e., the tool loads all scans overlapping with the current tile location.
Thus, you will always label the part of the scans that overlaps with the current tile.
In the `settings.cfg` files you can change the followings options:
Given the poses of a KITTI point cloud dataset, we load tiles of overlapping point clouds. Thus, multiple point clouds are labeled at once in a certain area.
## Features
- Support for KITTI Vision Benchmark Point Clouds.
- Human-readable label description files in xml allow to define label names, ids, and colors.
- Modern OpenGL shaders for rendering of even millions of points.
- Tools for labeling of individual points and polygons.
- Filtering of labels makes it easy to label even complicated structures with ease.
## Dependencies
* catkin
* Eigen >= 3.2
* boost >= 1.54
* QT >= 5.2
* OpenGL Core Profile >= 4.0
* [glow](https://github.com/jbehley/glow) (catkin package)
## Build
On Ubuntu 16.04 and 18.04, most of the dependencies can be installed from the package manager:
```bash
sudo apt install git libeigen3-dev libboost-all-dev qtbase5-dev libglew-dev catkin
```
Additionally, make sure you have [catkin-tools](https://catkin-tools.readthedocs.io/en/latest/) and the [fetch](https://github.com/Photogrammetry-Robotics-Bonn/catkin_tools_fetch) verb installed:
```bash
sudo apt install python-pip
sudo pip install catkin_tools catkin_tools_fetch empy
```
If you do not have a catkin workspace already, create one:
```bash
cd
mkdir catkin_ws
cd catkin_ws
mkdir src
catkin init
cd src
git clone https://github.com/ros/catkin.git
```
Clone the repository in your catkin workspace:
```bash
cd ~/catkin_ws/src
git clone https://github.com/jbehley/point_labeler.git
```
Download the additional dependencies:
```bash
catkin deps fetch
```
Then, build the project:
```bash
catkin build point_labeler
```
Now the project root directory (e.g. `~/catkin_ws/src/point_labeler`) should contain a `bin` directory containing the labeler.
## Usage
In the `bin` directory, just run `./labeler` to start the labeling tool.
The labeling tool allows to label a sequence of point clouds in a tile-based fashion, i.e., the tool loads all scans overlapping with the current tile location.
Thus, you will always label the part of the scans that overlaps with the current tile.
In the `settings.cfg` files you can change the followings options:
tile size: 100.0 # size of a tile (the smaller the less scans get loaded.)
max scans: 500 # number of scans to load for a tile. (should be maybe 1000), but this currently very memory consuming.
min range: 0.0 # minimum distance of points to consider.
max range: 50.0 # maximum distance of points in the point cloud.
## Folder structure
When loading a dataset, the data must be organized as follows:
point cloud folder
├── velodyne/ -- directory containing ".bin" files with Velodyne point clouds.
├── labels/ [optional] -- label directory, will be generated if not present.
├── image_2/ [optional] -- directory containing ".png" files from the color camera.
├── calib.txt -- calibration of velodyne vs. camera. needed for projection of point cloud into camera.
└── poses.txt -- file containing the poses of every scan.
## Documentation
See the [wiki](https://github.com/jbehley/point_labeler/wiki) for more information on the usage and other details.
## Citation
If you're using the tool in your research, it would be nice if you cite our [paper](https://arxiv.org/abs/1904.01416):
```
@inproceedings{behley2019iccv,
author = {J. Behley and M. Garbade and A. Milioto and J. Quenzel and S. Behnke and C. Stachniss and J. Gall},
title = {{SemanticKITTI: A Dataset for Semantic Scene Understanding of LiDAR Sequences}},
booktitle = {Proc. of the IEEE/CVF International Conf.~on Computer Vision (ICCV)},
year = {2019}
}
```
We used the tool to label SemanticKITTI, which contains overall over 40.000 scans organized in 20 sequences.  Given the poses of a KITTI point cloud dataset, we load tiles of overlapping point clouds. Thus, multiple point clouds are labeled at once in a certain area.
## Features
- Support for KITTI Vision Benchmark Point Clouds.
- Human-readable label description files in xml allow to define label names, ids, and colors.
- Modern OpenGL shaders for rendering of even millions of points.
- Tools for labeling of individual points and polygons.
- Filtering of labels makes it easy to label even complicated structures with ease.
## Dependencies
* catkin
* Eigen >= 3.2
* boost >= 1.54
* QT >= 5.2
* OpenGL Core Profile >= 4.0
* [glow](https://github.com/jbehley/glow) (catkin package)
## Build
On Ubuntu 16.04 and 18.04, most of the dependencies can be installed from the package manager:
```bash
sudo apt install git libeigen3-dev libboost-all-dev qtbase5-dev libglew-dev catkin
```
Additionally, make sure you have [catkin-tools](https://catkin-tools.readthedocs.io/en/latest/) and the [fetch](https://github.com/Photogrammetry-Robotics-Bonn/catkin_tools_fetch) verb installed:
```bash
sudo apt install python-pip
sudo pip install catkin_tools catkin_tools_fetch empy
```
If you do not have a catkin workspace already, create one:
```bash
cd
mkdir catkin_ws
cd catkin_ws
mkdir src
catkin init
cd src
git clone https://github.com/ros/catkin.git
```
Clone the repository in your catkin workspace:
```bash
cd ~/catkin_ws/src
git clone https://github.com/jbehley/point_labeler.git
```
Download the additional dependencies:
```bash
catkin deps fetch
```
Then, build the project:
```bash
catkin build point_labeler
```
Now the project root directory (e.g. `~/catkin_ws/src/point_labeler`) should contain a `bin` directory containing the labeler.
## Usage
In the `bin` directory, just run `./labeler` to start the labeling tool.
The labeling tool allows to label a sequence of point clouds in a tile-based fashion, i.e., the tool loads all scans overlapping with the current tile location.
Thus, you will always label the part of the scans that overlaps with the current tile.
In the `settings.cfg` files you can change the followings options:
Given the poses of a KITTI point cloud dataset, we load tiles of overlapping point clouds. Thus, multiple point clouds are labeled at once in a certain area.
## Features
- Support for KITTI Vision Benchmark Point Clouds.
- Human-readable label description files in xml allow to define label names, ids, and colors.
- Modern OpenGL shaders for rendering of even millions of points.
- Tools for labeling of individual points and polygons.
- Filtering of labels makes it easy to label even complicated structures with ease.
## Dependencies
* catkin
* Eigen >= 3.2
* boost >= 1.54
* QT >= 5.2
* OpenGL Core Profile >= 4.0
* [glow](https://github.com/jbehley/glow) (catkin package)
## Build
On Ubuntu 16.04 and 18.04, most of the dependencies can be installed from the package manager:
```bash
sudo apt install git libeigen3-dev libboost-all-dev qtbase5-dev libglew-dev catkin
```
Additionally, make sure you have [catkin-tools](https://catkin-tools.readthedocs.io/en/latest/) and the [fetch](https://github.com/Photogrammetry-Robotics-Bonn/catkin_tools_fetch) verb installed:
```bash
sudo apt install python-pip
sudo pip install catkin_tools catkin_tools_fetch empy
```
If you do not have a catkin workspace already, create one:
```bash
cd
mkdir catkin_ws
cd catkin_ws
mkdir src
catkin init
cd src
git clone https://github.com/ros/catkin.git
```
Clone the repository in your catkin workspace:
```bash
cd ~/catkin_ws/src
git clone https://github.com/jbehley/point_labeler.git
```
Download the additional dependencies:
```bash
catkin deps fetch
```
Then, build the project:
```bash
catkin build point_labeler
```
Now the project root directory (e.g. `~/catkin_ws/src/point_labeler`) should contain a `bin` directory containing the labeler.
## Usage
In the `bin` directory, just run `./labeler` to start the labeling tool.
The labeling tool allows to label a sequence of point clouds in a tile-based fashion, i.e., the tool loads all scans overlapping with the current tile location.
Thus, you will always label the part of the scans that overlaps with the current tile.
In the `settings.cfg` files you can change the followings options: