
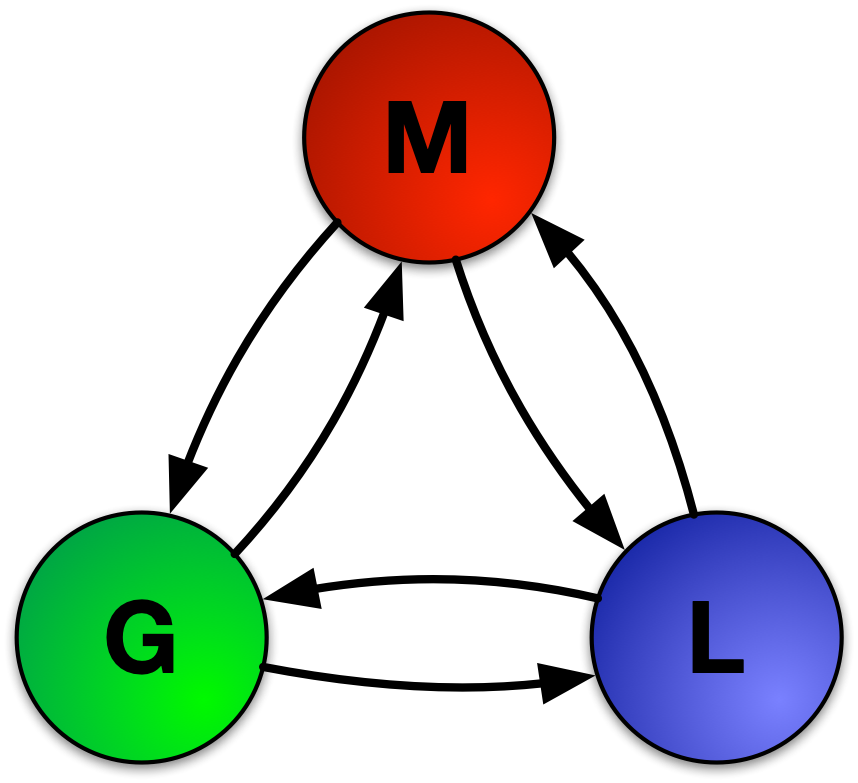
Figure: Schematic of M3GNet/MEGNet
Here, we summarize the currently implemented architectures in MatGL. It should be stressed that this is by no means
an exhaustive list, and we expect new architectures to be added by the core MatGL team as well as other contributors
in the future.
- [QET] (DGL only, PYG coming soon), pronounced as "ket", is a charge-equilibrated TensorNet architecture. It is an
equivariant, charge-aware architecture that attains linear scaling with system size via an analytically solvable
charge-equilibration scheme. A pre-trained QET-MatQ FP is available, which matches state-of-the-art FPs on standard
materials property benchmarks but delivers qualitatively different predictions in systems dominated by charge
transfer, e.g., NaCl–\ce{CaCl2} ionic liquid, reactive processes at the Li/\ce{Li6PS5Cl} solid-electrolyte interface,
and supports simulations under applied electrochemical potentials.
- [TensorNet] (PYG and DGL) is an O(3)-equivariant message-passing neural network architecture that leverages Cartesian tensor
representations. It is a generalization of the [SO3Net] architecture, which is a minimalist SO(3)-equivariant neural
network. In general, TensorNet has been shown to be much more data and parameter efficient than other equivariant
architectures. It is currently the default architecture used in the [Materials Virtual Lab].
- [Crystal Hamiltonian Graph Network (CHGNet)][chgnet] (DGL only) is a graph neural network based MLIP. CHGNet involves atom
graphs to capture atom bond relations and bond graph to capture angular information. It specializes in
capturing the atomic charges through learning and predicting DFT atomic magnetic moments.
See [original implementation][chgnetrepo]
- [Materials 3-body Graph Network (M3GNet)][m3gnet] is an invariant graph neural network architecture that
incorporates 3-body interactions. An additional difference is the addition of the coordinates for atoms and
the 3×3 lattice matrix in crystals, which are necessary for obtaining tensorial quantities such as forces and
stresses via auto-differentiation. As a framework, M3GNet has diverse applications, including **Interatomic potential development.** With the same training data, M3GNet performs similarly to state-of-the-art
machine learning interatomic potentials (MLIPs). However, a key feature of a graph representation is its
flexibility to scale to diverse chemical spaces. One of the key accomplishments of M3GNet is the development of a
[*foundation potential*][m3gnet] that can work across the entire periodic table of the elements by training on
relaxations performed in the [Materials Project][mp]. Like the previous MEGNet architecture, M3GNet can be used to
develop surrogate models for property predictions, achieving in many cases accuracies that are better or similar to
other state-of-the-art ML models.
- [MatErials Graph Network (MEGNet)][megnet] (DGL only) is an implementation of DeepMind's [graph networks][graphnetwork] for
machine learning in materials science. We have demonstrated its success in achieving low prediction errors in a broad
array of properties in both [molecules and crystals][megnet]. New releases have included our recent work on
[multi-fidelity materials property modeling][mfimegnet]. Figure 1 shows the sequential update steps of the graph
network, whereby bonds, atoms, and global state attributes are updated using information from each other, generating
an output graph.
For detailed performance benchmarks, please refer to the publications in the [References](#references) section.
## Installation
Matgl can be installed via pip:
```bash
pip install matgl
```
If you need to use DGL, it is recommended you install the latest version of DGL before installing matgl.
```bash
pip install dgl -f https://data.dgl.ai/wheels/torch-2.4/repo.html
```
### CUDA (GPU) installation
If you intend to use CUDA (GPU) to speed up training, it is important to install the appropriate versions of PyTorch
and DGL. The basic instructions are given below, but it is recommended that you consult the
[PyTorch docs](https://pytorch.org/get-started/locally/) and [DGL docs](https://www.dgl.ai/pages/start.html) if you
run into any problems.
```shell
pip install torch==2.2.0 --index-url https://download.pytorch.org/whl/cu121
pip install dgl -f https://data.dgl.ai/wheels/cu121/repo.html
pip install dglgo -f https://data.dgl.ai/wheels-test/repo.html
```
## Docker images
Docker images have now been built for matgl, together with LAMMPS support. They are available at the
[Materials Virtual Lab Docker Repository]. If you wish to use MatGL with LAMMPS, this is probably the easiest option.
## Usage
Pre-trained M3GNet universal potential and MEGNet models for the Materials Project formation energy and
multi-fidelity band gap are now available.
### Command line (from v0.6.2)
A CLI tool now provides the capability to perform quick relaxations or predictions using pre-trained models, as well
as other simple administrative tasks (e.g., clearing the cache). Some simple examples:
1. To perform a relaxation,
```bash
mgl relax --infile Li2O.cif --outfile Li2O_relax.cif
```
2. To use one of the pre-trained property models,
```bash
mgl predict --model M3GNet-MP-2018.6.1-Eform --infile Li2O.cif
```
3. To clear the cache,
```bash
mgl clear
```
For a full range of options, use `mgl -h`.
### Code
Users who just want to use the models out of the box should use the newly implemented `matgl.load_model` convenience
method. The following is an example of a prediction of the formation energy for CsCl.
```python
from pymatgen.core import Lattice, Structure
import matgl
model = matgl.load_model("MEGNet-MP-2018.6.1-Eform")
# This is the structure obtained from the Materials Project.
struct = Structure.from_spacegroup("Pm-3m", Lattice.cubic(4.1437), ["Cs", "Cl"], [[0, 0, 0], [0.5, 0.5, 0.5]])
eform = model.predict_structure(struct)
print(f"The predicted formation energy for CsCl is {float(eform.numpy()):.3f} eV/atom.")
```
To obtain a listing of available pre-trained models,
```python
import matgl
print(matgl.get_available_pretrained_models())
```
## Pytorch Hub
The pre-trained models are also available on Pytorch hub. To use these models, simply install matgl and use the
following commands:
```python
import torch
# To obtain a listing of models
torch.hub.list("materialsvirtuallab/matgl", force_reload=True)
# To load a model
model = torch.hub.load("materialsvirtuallab/matgl", 'm3gnet_universal_potential')
```
## Model Training
In the PES training, the unit of energies, forces and stresses (optional) in the training, validation and test sets is extremely important to be consistent with the unit used in MatGL.
- energies: a list of energies with unit eV.
- forces: a list of nx3 force matrix with unit eV/Å, where n is the number of atom in each structure. n does not need to be the same for all structures.
- stresses: a list of 3x3 stress matrices with unit GPa (optional)
Note: For stresses, we use the convention that compressive stress gives negative values. Stresses obtained from VASP calculations (default unit is kBar) should be multiplied by -0.1 to work directly with the model.
## Tutorials
We wrote [tutorials] on how to use MatGL. These were generated from [Jupyter notebooks]
[jupyternb], which can be directly run on [Google Colab].
## Resources
- [API docs][apidocs] for all classes and methods.
- [Developer Guide](developer.md) outlines the key design elements of `matgl`, especially for developers wishing to
train and contribute matgl models.
- AdvancedSoft has implemented a [LAMMPS interface](https://github.com/advancesoftcorp/lammps/tree/based-on-lammps_2Jun2022/src/ML-M3GNET)
to both the TF and MatGL version of M3GNet.
## References
A manuscript for MatGL has been published in npj Computational Materials. please cite the following:
> **MatGL**
>
> Ko, T. W.; Deng, B.; Nassar, M.; Barroso-Luque, L.; Liu, R.; Qi, J.; Thakur, A. C.; Mishra, A. R.; Liu, E.; Ceder, G.; Miret, S.; Ong, S. P.
> *Materials Graph Library (MatGL), an Open-Source Graph Deep Learning Library for Materials Science and Chemistry.*
> npj Comput Mater 11, 253 (2025). DOI: [https://doi.org/10.1038/s41524-025-01742-y][matgl].
If you are using any of the pretrained models, please cite the relevant works below:
> **MEGNet**
>
> Chen, C.; Ye, W.; Zuo, Y.; Zheng, C.; Ong, S. P. *Graph Networks as a Universal Machine Learning Framework for
> Molecules and Crystals.* Chem. Mater. 2019, 31 (9), 3564–3572. DOI: [10.1021/acs.chemmater.9b01294][megnet].
> **Multi-fidelity MEGNet**
>
> Chen, C.; Zuo, Y.; Ye, W.; Li, X.; Ong, S. P. *Learning Properties of Ordered and Disordered Materials from
> Multi-Fidelity Data.* Nature Computational Science, 2021, 1, 46–53. DOI: [10.1038/s43588-020-00002-x][mfimegnet].
> **M3GNet**
>
> Chen, C., Ong, S.P. *A universal graph deep learning interatomic potential for the periodic table.* Nature
> Computational Science, 2023, 2, 718–728. DOI: [10.1038/s43588-022-00349-3][m3gnet].
>**CHGNet**
>
> Deng, B., Zhong, P., Jun, K. et al. *CHGNet: as a pretrained universal neural network potential for charge-informed atomistic modelling.*
> Nat Mach Intell 5, 1031–1041 (2023). DOI:[10.1038/s42256-023-00716-3][chgnet]
>**TensorNet**
>
> Simeon, G. De Fabritiis, G. *Tensornet: Cartesian tensor representations for efficient learning of molecular potentials.*
> Adv. Neural Info. Process. Syst. 36, (2024). DOI: [10.48550/arXiv.2306.06482][tensornet]
>**SO3Net**
>
> Schütt, K. T., Hessmann, S. S. P., Gebauer, N. W. A., Lederer, J., Gastegger, M. *SchNetPack 2.0: A neural network toolbox for atomistic machine learning.*
> J. Chem. Phys. 158, 144801 (2023). DOI: [10.1063/5.0138367][so3net]
## FAQs
1. **The `M3GNet-MP-2021.2.8-PES` differs from the original TensorFlow (TF) implementation!**
*Answer:* `M3GNet-MP-2021.2.8-PES` is a refitted model with some data improvements and minor architectural changes.
Porting over the weights from the TF version to DGL/PyTorch is non-trivial. We have performed reasonable benchmarking
to ensure that the new implementation reproduces the broad error characteristics of the original TF implementation
(see [examples][jupyternb]). However, it is not expected to reproduce the TF version exactly. This refitted model
serves as a baseline for future model improvements. We do not believe there is value in expending the resources
to reproduce the TF version exactly.
2. **I am getting errors with `matgl.load_model()`!**
*Answer:* The most likely reason is that you have a cached older version of the model. We often refactor models to
ensure the best implementation. This can usually be solved by updating your `matgl` to the latest version
and clearing your cache using the following command `mgl clear`. On the next run, the latest model will be
downloaded. With effect from v0.5.2, we have implemented a model versioning scheme that will detect code vs model
version conflicts and alert the user of such problems.
3. **What pre-trained models should I be using?**
*Answer:* There is no one definitive answer. In general, the newer the architecture and dataset, the more likely
the model performs better. However, it should also be noted that a model operating on a more diverse dataset may
compromise on performance on a specific system. The best way is to look at the READMEs included with each model
and do some tests on the systems you are interested in.
4. **How do I contribute to matgl?**
*Answer:* For code contributions, please fork and submit pull requests. You should read the
[developer guide](developer.md) to understand the general design guidelines. We welcome pre-trained model
contributions as well, which should also be submitted via PRs. Please follow the folder structure of the
pretrained models. In particular, we expect all models to come with a `README.md` and notebook
documenting its use and its key performance metrics. Also, we expect contributions to be on new properties
or systems or to significantly outperform the existing models. We will develop an alternative means for model
sharing in the future.
5. **None of your models do what I need. Where can I get help?**
*Answer:* Please contact [Prof Ong][ongemail] with a brief description of your needs. For simple problems, we are
glad to advise and point you in the right direction. For more complicated problems, we are always open to
academic collaborations or projects. We also offer [consulting services][mqm] for companies with unique needs,
including but not limited to custom data generation, model development and materials design.
## Acknowledgments
This work was primarily supported by the [Materials Project][mp], funded by the U.S. Department of Energy, Office of
Science, Office of Basic Energy Sciences, Materials Sciences and Engineering Division under contract no.
DE-AC02-05-CH11231: Materials Project program KC23MP. This work used the Expanse supercomputing cluster at the Extreme
Science and Engineering Discovery Environment (XSEDE), which is supported by National Science Foundation grant number
ACI-1548562.
[m3gnetrepo]: https://github.com/materialsvirtuallab/m3gnet "M3GNet repo"
[megnetrepo]: https://github.com/materialsvirtuallab/megnet "MEGNet repo"
[dgl]: https://www.dgl.ai "DGL website"
[mavrl]: http://materialsvirtuallab.org "MAVRL website"
[changelog]: https://matgl.ai/changes "Changelog"
[graphnetwork]: https://arxiv.org/abs/1806.01261 "Deepmind's paper"
[megnet]: https://pubs.acs.org/doi/10.1021/acs.chemmater.9b01294 "MEGNet paper"
[mfimegnet]: https://nature.com/articles/s43588-020-00002-x "mfi MEGNet paper"
[m3gnet]: https://nature.com/articles/s43588-022-00349-3 "M3GNet paper"
[mp]: http://materialsproject.org "Materials Project"
[apidocs]: https://matgl.ai/matgl.html "MatGL API docs"
[doc]: https://matgl.ai "MatGL Documentation"
[google colab]: https://colab.research.google.com/ "Google Colab"
[jupyternb]: https://github.com/materialsvirtuallab/matgl/tree/main/examples
[ongemail]: mailto:ongsp@ucsd.edu "Email"
[mqm]: https://materialsqm.com "MaterialsQM"
[tutorials]: https://matgl.ai/tutorials "Tutorials"
[matgl]: https://www.nature.com/articles/s41524-025-01742-y#citeas "MatGL"
[tensornet]: https://arxiv.org/abs/2306.06482 "TensorNet"
[qet]: https://arxiv.org/abs/2511.07249 "QET"
[so3net]: https://pubs.aip.org/aip/jcp/article-abstract/158/14/144801/2877924/SchNetPack-2-0-A-neural-network-toolbox-for "SO3Net"
[chgnet]: https://www.nature.com/articles/s42256-023-00716-3 "CHGNet"
[chgnetrepo]: https://github.com/CederGroupHub/chgnet "CHGNet repo"
[maml]: https://materialsvirtuallab.github.io/maml/
[MatGL]: https://matgl.ai
[MatPES]: https://matpes.ai
[MatCalc]: https://matcalc.ai
[Materials Virtual Lab Docker Repository]: https://hub.docker.com/orgs/materialsvirtuallab/repositories
 ## Official Documentation
## Official Documentation
